Introduction to Crossmaps
A (Nomenclature) Crossmap encodes a complete redistribution of values between a source and target classifications as a directed, bipartite, weighted graph. Numeric values transformed using a valid Crossmap will sum to the same total in both the source and target classifications. A valid Crossmap satisfies the following conditions:
- There is at most one link between each distinct source and target node
- For each source node, the sum of weights attached to all outgoing links sums to one.
For example, this mapping between occupation codes in ANZSCO22 and ISCO8 is a valid crossmap:
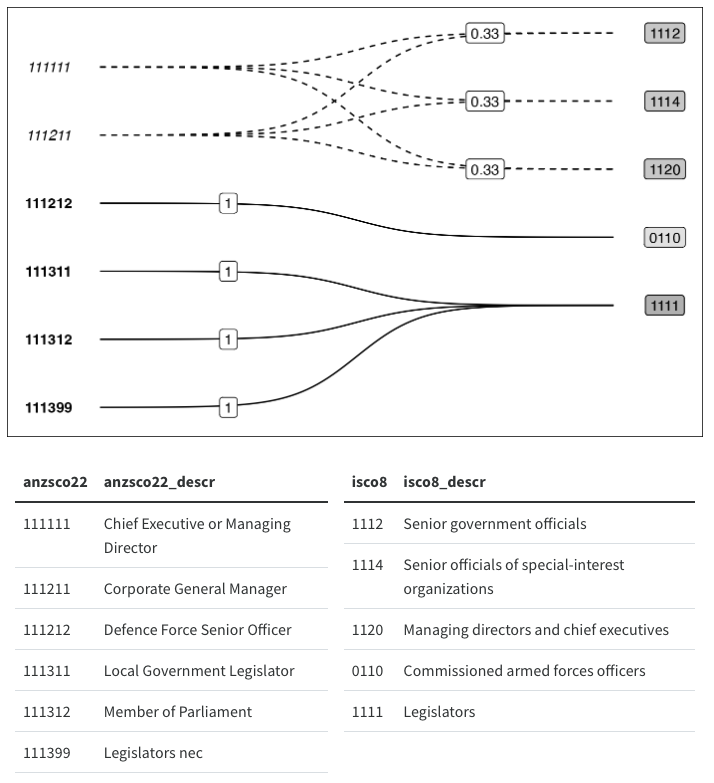
Unit and Fractional Weight Links
The weights associated with each source-target link encode the transformation to be applied to values associated with a given source node. Let the weight between source node \(i\) and target node \(j\) be denoted \(w_{ij}\). Weights can range from \([0,1]\). Note that \(w_{ij} = 0\) is the trivial case where there is no link between the source and target nodes. The non-trivial cases are unit weights, \(w_{ij} = 1\) and fractional weights, \(w_{ij} \in (0,1)\).
Unit weights indicate that source values will be unmodified when passed into the target nomenclature. Therefore, crosswalks for recoding node labels, or collapsing multiple sub-category source nodes into parent target nodes, can both be represented as crossmaps with unit weights.
Fractional weights on the other hand indicate how a source value will be modified when passed to the target node. For example a link with a weight of \(0.7\) indicates that 70% of the numeric value associated with a source node should be “passed over” to the target node.
Now, consider two links from the same source node \(i\) to two different target nodes \(j\) and \(k\), with weights \(w_{ij}\) and \(w_{ik}\). Assume that the source node \(i\) has no other outgoing links into the target nomenclature. Then, it follows that the weights on the two given links should sum to one if we want to preserve the total value between the source and target nomenclature. For instance, if \(w_{ij} = 0.7\), then \(w_{ik}\) should be \(0.3\), such that 70% of the value associated with source node \(i\) goes to the target node \(j\), and the remaining 30% is distributed to target node \(k\).
Crossmap tables
Crossmap tables are extensions of crosswalk or lookup tables, where
each row represents a weighted link between the source and target
classifications. You can pass a data.frame or tibble of weighted edges
to verify_links_as_xmap() to check if the input satisfies
crossmap properties.
Your input table x needs to have at least 3 complete
(i.e. no NA) columns:
-
from: source classification labels. -
to: target classification labels -
weights: applied to values in the source classification
Additional columns (e.g. for label descriptions etc.) can be retained
by setting the .drop_extra argument to
FALSE.
verify_links_as_xmap and as_xmap_df() will
validate that:
- There are no duplicates
from-topairs, - Every group of
weightsassociated with a distinctfromvalue sums to 1 (subject to a floating point tolerance).
The package also offers a experimental xmap_df class
facilitate additional functionality such as graph property calculation
and printing (i.e. relation types and crossmap direction) via a custom
print() method, coercion to other useful classes
(e.g. xmap_to_matrix()), visualisation (see
vignette("vis-xmaps") for prototypes), and multi-step
transformations (i.e. from nomenclature A to B to C).
Please note that the xmap_df class and related functions
are still in active development and subject to breaking changes in
future releases.
Special Cases of Crossmaps
The conditions for valid nomenclature crossmaps defined above are sufficient to ensure that the total sum of value is retained between the source and target nomenclature However, there are a number of other transformation properties which crossmaps can also encode and validate. Such special cases include “degenerate” crossmaps. We consider a crossmap “degenerate” if it has only binary weights on all possible source-target links. In such cases the weights are implied by the presence or absence of a link.
Types of Mapping Relations
It is helpful when thinking about special cases to define some familiar types of relations:
- one-to-one: a set of links where pairs of source and target nodes are uniquely linked to each other and no other nodes. Any links in this type of relation will have unit weights.
- one-to-many: a set of links where a single source node is linked to multiple target nodes. Links in this “splitting” relation will have fractional weights.
- one-from-many: a set of links where a single target node is linked to multiple source nodes. This “collapse” relation is more commonly known as a many-to-one. Links can have either unit or fractional weights depending on whether the source nodes are part of one-to-one or one-to-many relations.
- many-to-many: refers to a combination of the above relations.
Note that links in a crossmap can be partitioned into subgraphs according to the number of outgoing links (i.e. one-to-one and one-to-many)
Recode maps
Crossmaps with only one-to-one relations recode source labels to target labels, leaving any attached values untouched. This implies that the weights on links are always 1. Notice that Schema Crosswalks, lookup tables and category recoding functions are all Recode Maps.
A common way of implementing recodings in R is named vectors or
pairwise arguments such as in forcats::fct_recode(). Named
vectors are a convenient way to store and use one-to-one and many-to-one
mappings. In general it is not necessary to convert such mappings into
crossmaps unless you want to combine them with fractional weight links.
However, when necessary, named vectors can be converted into
xmap by first transforming vector into two-column table of
node pairs, and then attaching the implied unit weights.
# example using the named vector included in `mock` objects:
mock$recode_vect
recode_xmap <-
xmap::as_pairs_from_named(mock$recode_vect, names_to = "iso3", values = "ctr_name") |>
xmap::add_weights_unit(weights_into = "w") |>
xmap::as_xmap_df(from = iso3, to = ctr_name, weights = w)For a crossmap to be a recode map it must have binary weights, and
the cardinality of the source and target node sets must be equal. In
other words, weights can only be 0 (i.e. no link) or 1, and the number
of unique source and target labels should be the same. The function
verify_named_all_1to1() or its alias
verify_named_as_recode_unique() uses the cardinality
condition to verify whether or not a named vector has only one-to-one
relation:
fruit_color <- c(apple = "green", strawberry = "red", banana = "yellow")
fruit_color |>
verify_named_all_1to1() |>
print()## apple strawberry banana
## "green" "red" "yellow"We can also identify mistakes such as accidentally assigning a value twice:
fruit_color_mistake <- c(fruit_color, pear = "green")
fruit_color_mistake |>
verify_named_as_recode_unique()## Error in verify_named_as_recode_unique(fruit_color_mistake): could not find function "verify_named_as_recode_unique"Collapse or Aggregation maps
Collapse crossmaps are similar to Recode crossmaps in that source values are unmodified when passed into the target nomenclature. However, as the name suggests, multiple source nodes can be “collapsed” into the same target node, such that at least some of the source values will be aggregated in the target nomenclature. Similar to a Recode crossmap, a Collapse crossmap must only have unit weights, but the cardinality of the source node set will be larger than the target node set. This means that at least two source nodes must be linked to the same target node since there are fewer target nodes than source nodes.
Similar to the recode maps, collapse maps only have unit weights. Hence, we can use named lists to store such mappings. Consider an example such as assigning students to groups. Assume no two students share the same name:
student_groups <- list(
GRP1 = c("kate", "jane", "peter"),
GRP2 = c("terry", "ben", "grace"),
GRP3 = c("cindy", "lucy", "alex")
)We can check that students are not assigned to multiple groups:
student_groups |>
verify_named_all_values_unique()Or that the every student has been assigned a group:
## mistakenly assign kate to another group
student_group_mistake <- list(
GRP1 = c("kate", "jane", "peter"),
GRP2 = c("terry", "ben", "grace"),
GRP2 = c("cindy", "lucy", "kate")
)
student_list <- c("kate", "jane", "peter", "terry", "ben", "grace", "cindy", "lucy", "alex")
student_group_mistake |>
verify_named_matchset_values_exact(student_list)## Error in `verify_named_matchset_values_exact()`:
## ! The values of `x` do not exactly match `ref_set`and each group has a unique name:
student_group_mistake |>
verify_named_all_names_unique()## Error in `verify_named_all_names_unique()`:
## ! Duplicated names found in `x`.
## ℹ Use `base::duplicated(names(x))` to identify duplicates.Coercion to xmap requires adding weights and is
directional. We can choose to verify conditions and return the same
object via verify_links_* functions, or coerce to an
xmap for use with other functions in the package.
group_links <- student_groups |>
as_pairs_from_named(names_to = "group", values_to = "student") |>
add_weights_unit()
## collapse xmap from students to groups
group_links |>
verify_links_as_xmap(from = student, to = group, weights = weights)
## reverse doesn't work without adjusting the weights
group_links |>
verify_links_as_xmap(from = group, to = student, weights = weights)## Error in `verify_links_as_xmap()`:
## ! Incomplete mapping weights found
## ✖ `weights` does not sum to 1
## ℹ Modify weights or adjust `tol` and try again.Aggregation transformations can implemented as Collapse maps. Additionally, aggregation size requirements can be implemented as conditions on the number of incoming links for each target node. For instance, if you know that each target node should aggregate values from at least two source nodes, then the minimum number of non-zero incoming links should be 2 for every target node. Notice that this requirement precludes one-to-one links since it prevents any source node from being recoded into an “unshared” node in the target nomenclature.
Split or Redistribution maps (with Equal Weights)
Any crossmap with at least one set of fractional weight (one-to-many) links involve some redistribution of source value. However, a useful special case of maps with fractional weights are split maps, which encode the disaggregation of source values into a target nomenclature. Such maps which will have mostly one-to-many relations, and do not contain one-from-many relations. This can be ensured by restricting each target node to only have one incoming link. Furthermore, we can preclude one-to-one links with the condition that all links must have fractional weights.
Notice that the same set of node pairs can generate either a split or
collapse map depending on the weights added. For example, recall that in
the student group example from above, swapping the to and
from variables no longer formed a valid crossmap. Now
consider allocating some budget of prizes equally between students in
each group. This requires adding fractional weights to
group_links, which can then be verified as a crossmap.
group_prize_links <- student_groups |>
as_pairs_from_named("group", "student") |>
dplyr::group_by(group) |>
dplyr::mutate(prize_share = 1 / dplyr::n_distinct(student))
group_prize_links |>
verify_links_as_xmap(from = group, to = student, weights = prize_share)Note that if the first crossmap condition is met, i.e. there is only
one link between each unique source-node pair, then
dplyr::n() and dplyr::n_distinct() are
interchangeable when generating equal share weights.
Cheatsheet for Verifying Mapping Objects
Let \(G = (U,V, E)\) be a crossmap where \(U\) and \(V\) are the source and target nomenclature sets, and \(E\) denotes the graph edges (i.e. links between the two sets), with weights on a link between node \(i \in U\) and \(j \in V\) denoted \(w_{ij}\). Recall also that sets by definition don’t contain duplicates.
Let us also define some corresponding R objects:
-
from,to,weights: the vectors or columns defining non-zero links for a crossmap (i.e an edge list). Each - triplet or row represents a edge \({i,j}\) with weight \(w_{ij}\). These vectors can contain duplicates. -
unique(from): the set \(U\) of unique categories in the source nomenclature -
unique(to): the set \(V\) of unique categories in the target nomenclature -
df: a data.frame containingfrom,to,weights -
sum_w: a vector the same length asunique(from)containing sum ofweightsgrouped byfrom -
ones: a vector of ones the same length assum_wandunique(from)
Then to check if table of links is a valid crossmap, we just check that:
- there are no missing values – i.e. any explicit link has to have a source, target and weight
- we only have one link between each unique source and target node – i.e. we can’t have two links between \(i,j\) with different weights.
- the weights on links coming out of each source node sum to 1
The xmap function to do this is
verify_links_as_xmap(). But you can also check the
following base R or dplyr conditions:
anyNA(df)-
anyDuplicated(data.frame(from, to)) == FALSEornrow(distinct(df, from, to)) = nrow(df) -
all(isTRUE(all.equal(sum_w, ones)))ordplyr::near(sum_w, ones)
Note that using == for the third condition is not
recommended since it doesn’t have any built-in tolerance for floating
point errors.
Strict versions of the above special cases can also be characterised with the following additional conditions:
| x | Recoding (1-to-1) | Collapsing (M-to-1) | Splitting (1-to-M) |
|---|---|---|---|
| Assertions |
|
|
|
| xmap functions |
… |
… |
… |
|
base R conditions |
|
|
|
| Graph Conditions |
|
|
|
Creating xmap_df from Pairs and Links
The following examples demonstrate how to coerce various inputs into
the experimental xmap_df class.
Row-wise Links
The following example shows how to create a mixed crossmap which
encodes one-to-one, many-to-one and one-to-many relations and coerce
them into a xmap_df. This method is most suitable for
simple crossmaps. The exmaple belows shows the code for the sample
crossmap: mock$xmap_abc
abc_links <- tibble::tribble(
~lower, ~upper, ~share,
"a", "AA", 1, # one-to-one
"b", "BB", 1, # one-FROM-many
"c", "BB", 1,
"d", "CC", 0.3, # one-to-many
"d", "DD", 0.6,
"d", "EE", 0.1
)
xmap_abc <- abc_links |>
as_xmap_df(from = lower, to = upper, weights = share)## Dropping any additional columns in `abc_links`
## ℹ To silence set `.drop_extra = TRUE`
xmap_abc## xmap_df:
## recode, split, and collapse
## (lower -> upper) BY share
## lower upper share
## 1 a AA 1.0
## 2 b BB 1.0
## 3 c BB 1.0
## 4 d CC 0.3
## 5 d DD 0.6
## 6 d EE 0.1Crosswalk/Lookup Tables
It is more common that you will want to convert an existing correspondence into a crossmap. Such conversions require attaching appropriate weights to the existing crosswalk table.
Recode Pairs
Consider the first five country codes in the ISO 3166 international standard and the one-to-one correspondence between the 2-digit, 2-digit and numeric codes.
iso_codes <- tibble::tribble(
~country, ~ISO2, ~ISO3, ~ISONumeric,
"Afghanistan", "AF", "AFG", "004",
"Albania", "AL", "ALB", "008",
"Algeria", "DZ", "DZA", "012",
"American Samoa", "AS", "ASM", "016",
"Andorra", "AD", "AND", "020"
)We can verify any pairwise combination of columns as one-to-one
recode maps using the verify_pairs_* functions:
iso_codes |>
verify_pairs_as_recode_unique(from = country, to = ISO2) |>
print()## # A tibble: 5 × 4
## country ISO2 ISO3 ISONumeric
## <chr> <chr> <chr> <chr>
## 1 Afghanistan AF AFG 004
## 2 Albania AL ALB 008
## 3 Algeria DZ DZA 012
## 4 American Samoa AS ASM 016
## 5 Andorra AD AND 020To create a crossmap between ISONumeric and
ISO2, we simply add a weights columns and coerce to
xmap_df:
iso_xmap <- iso_codes |>
add_weights_unit(weights_into = "weight") |>
as_xmap_df(from = ISONumeric, to = ISO2, weights = weight)## Dropping any additional columns in `add_weights_unit(iso_codes, weights_into =
## "weight")`
## ℹ To silence set `.drop_extra = TRUE`Notice that as_xmap_df() always place the
from, to and weights columns in
order and drops any additional columns passed to it.
print(iso_xmap)## xmap_df:
## recode
## (ISONumeric -> ISO2) BY weight
## ISONumeric ISO2 weight
## 1 004 AF 1
## 2 008 AL 1
## 3 012 DZ 1
## 4 016 AS 1
## 5 020 AD 1To convert the validated xmap into a named vector:
iso_xmap |>
xmap_to_named_vector()## AF AL DZ AS AD
## "004" "008" "012" "016" "020"We can also easily generate validated crossmaps for the other nomenclature in the table, and keep additional columns:
iso_codes |>
add_weights_unit(weights_into = "weight") |>
as_xmap_df(from = ISO2, to = ISO3, weights = weight, .drop_extra = FALSE)## xmap_df:
## recode
## (ISO2 -> ISO3) BY weight
## ISO2 ISO3 weight country ISONumeric
## 1 AF AFG 1 Afghanistan 004
## 2 AL ALB 1 Albania 008
## 3 DZ DZA 1 Algeria 012
## 4 AS ASM 1 American Samoa 016
## 5 AD AND 1 Andorra 020Nested Lists
Now consider aggregating data which were collected using the ISO 3166-2 Subdivisions of Australia and Canada:
adm1_list <- tibble::tribble(
~ctr, ~adm1,
"AU", "AU-NSW, AU-QLD, AU-SA, AU-TAS, AU-VIC, AU-WA, AU-ACT, AU-NT",
"CA", "CA-AB, CA-BC, CA-MB, CA-NB, CA-NL, CA-NS, CA-ON, CA-PE, CA-QC, CA-SK, CA-NT, CA-NU, CA-YT"
)Recall that we need one row per relation between the source
(adm1) and target (ctr) nomenclature. Thus we
split the string list into a vector, and then unnest the values by
country.
agg_x <- adm1_list |>
dplyr::mutate(adm1 = stringr::str_split(adm1, ", ")) |>
tidyr::unnest(cols = c(adm1))
agg_x## # A tibble: 21 × 2
## ctr adm1
## <chr> <chr>
## 1 AU AU-NSW
## 2 AU AU-QLD
## 3 AU AU-SA
## 4 AU AU-TAS
## 5 AU AU-VIC
## 6 AU AU-WA
## 7 AU AU-ACT
## 8 AU AU-NT
## 9 CA CA-AB
## 10 CA CA-BC
## # ℹ 11 more rowsSince aggregation involves the one-to-one transfer of values between
adm1 and ctr prior to the collapsing the
ctr groups, we simple add unit weights to form a valid
crossmap:
agg_xmap <- agg_x |>
add_weights_unit(weights_into = "link") |>
as_xmap_df(from = adm1, to = ctr, weights = link)## Dropping any additional columns in `add_weights_unit(agg_x, weights_into =
## "link")`
## ℹ To silence set `.drop_extra = TRUE`
agg_xmap## xmap_df:
## recode and collapse
## (adm1 -> ctr) BY link
## adm1 ctr link
## 1 AU-NSW AU 1
## 2 AU-QLD AU 1
## 3 AU-SA AU 1
## 4 AU-TAS AU 1
## 5 AU-VIC AU 1
## 6 AU-WA AU 1
## 7 AU-ACT AU 1
## 8 AU-NT AU 1
## 9 CA-AB CA 1
## 10 CA-BC CA 1
## 11 CA-MB CA 1
## 12 CA-NB CA 1
## 13 CA-NL CA 1
## 14 CA-NS CA 1
## 15 CA-ON CA 1
## 16 CA-PE CA 1
## 17 CA-QC CA 1
## 18 CA-SK CA 1
## 19 CA-NT CA 1
## 20 CA-NU CA 1
## 21 CA-YT CA 1References for Custom Weights
Conversely, we might have aggregate level data which want to disaggregate. Continuing the above example, this could involve incorporating country level data into analysis at the 3166-2 Subdivisions level.
For example, imagine that we have population figures for Australia at the 3166-2 level for 9 out of 10 years, but only country level figures for the missing year. In this simple example, a reasonable harmonisation design could involve splitting the country level figure by the subdivision level population proportions from the year preceding or following the missing year.
Alternatively, we might want to compare aggregate and disaggregate statistics to identify discrepancies.
Using Australian population statistics from 30 Jun 20221
state_data <- tibble::tribble(
~state, ~adm1, ~Pop,
"New South Wales", "AU-NSW", 8153600,
"Victoria", "AU-VIC", 6613700,
"Queensland", "AU-QLD", 5322100,
"South Australia", "AU-SA", 1820500,
"Western Australia", "AU-WA", 2785300,
"Tasmania", "AU-TAS", 571500,
"Northern Territory", "AU-NT", 250600,
"Australian Capital Territory", "AU-ACT", 456700
)
state_xmap <- state_data |>
dplyr::mutate(
ctr = "AU",
adm1,
share = Pop / sum(Pop)
) |>
as_xmap_df(from = ctr, to = adm1, weights = share)## Dropping any additional columns in `dplyr::mutate(state_data, ctr = "AU", adm1,
## share = Pop/sum(Pop))`
## ℹ To silence set `.drop_extra = TRUE`
state_xmap## xmap_df:
## split
## (ctr -> adm1) BY share
## ctr adm1 share
## 1 AU AU-NSW 0.31391391
## 2 AU AU-VIC 0.25462770
## 3 AU AU-QLD 0.20490105
## 4 AU AU-SA 0.07008932
## 5 AU AU-WA 0.10723416
## 6 AU AU-TAS 0.02200277
## 7 AU AU-NT 0.00964811
## 8 AU AU-ACT 0.01758297Piecewise Construction
Now consider the following mixed transformation using selected correspondences between NAICS Canada 1997 and ISIC Revision 3. Imagine that we have some numeric data (e.g. gross output in CAD) collected in the NAICS Canada nomenclature that we want to harmonise into ISIC Revision 3. The correspondence between the two nomenclature contains a mixture of one-to-one, one-to-many, and one-from-many relations.
Luckily, we can split the source nomenclature into two groups:
- source nodes with only one outgoing link (one-to-one relations with unit weights)
- source nodes with multiple outgoing links (one-to-many relations with fractional weights)
Let’s first define somed example correspondences.2
In the first example, one NAICS Canada class relates to exactly one ISIC class, forming a one-to-one relation.
canada_recode <- tibble::tibble(
NAICS1997 = "212210",
NAICS1997_desc = "Iron Ore Mining",
ISIC3 = "C1310",
ISIC3_desc = "Mining of iron ores"
)| NAICS1997 | NAICS1997_desc | ISIC3 | ISIC3_desc |
|---|---|---|---|
| 212210 | Iron Ore Mining | C1310 | Mining of iron ores |
In the second example, the ISIC target class D1543 is
equivalent to more than one NAICS Canada source class, forming a
one-from-many relation. The asterisk (Partial Flag) indicates that part
of ISIC D1543 is equivalent to each NAICS Canada class. The ISIC
activities corresponding to each NAICS Canada class are listed in the
column labelled “Link”.
canada_agg <- tibble::tribble(
~NAICS1997, ~NAICS1997_desc, ~ISIC3, ~ISIC3_desc, ~Link,
"311320", "Chocolate and Confectionery Manufacturing from Cacao Beans", "D1543 *", "Manufacture of cocoa, chocolate and sugar confectionery", "Chocolate and confectionery, made from cacao beans",
"311330", "Confectionery Manufacturing from Purchased Chocolate", "D1543 *", "Manufacture of cocoa, chocolate and sugar confectionery", "Confectionery, made from purchased chocolate",
"311340", "Non-Chocolate Confectionery Manufacturing", "D1543 *", "Manufacture of cocoa, chocolate and sugar confectionery", "Non-chocolate confectionery, manufacturing"
)| NAICS1997 | NAICS1997_desc | ISIC3 | ISIC3_desc | Link |
|---|---|---|---|---|
| 311320 | Chocolate and Confectionery Manufacturing from Cacao Beans | D1543 * | Manufacture of cocoa, chocolate and sugar confectionery | Chocolate and confectionery, made from cacao beans |
| 311330 | Confectionery Manufacturing from Purchased Chocolate | D1543 * | Manufacture of cocoa, chocolate and sugar confectionery | Confectionery, made from purchased chocolate |
| 311340 | Non-Chocolate Confectionery Manufacturing | D1543 * | Manufacture of cocoa, chocolate and sugar confectionery | Non-chocolate confectionery, manufacturing |
Notice that for both one-to-one and one-from-many relations, values attached to each source category are not directly modified during the “transfer” between source and target nomenclature. Instead, the source values are either retained, or summarised when the category collapse (and value aggregation) is performed. Thus, as shown above, all links in the above examples will take unit weights.
Let’s clean up the recode shown above:
canada_unit <- canada_agg |>
# remove the partial flag (*)
dplyr::mutate(ISIC3 = stringr::str_remove(ISIC3, " \\*")) |>
dplyr::select(-Link) |>
# bind the links together and add weights
dplyr::bind_rows(canada_recode) |>
dplyr::mutate(share = 1)| NAICS1997 | NAICS1997_desc | ISIC3 | ISIC3_desc | share |
|---|---|---|---|---|
| 311320 | Chocolate and Confectionery Manufacturing from Cacao Beans | D1543 | Manufacture of cocoa, chocolate and sugar confectionery | 1 |
| 311330 | Confectionery Manufacturing from Purchased Chocolate | D1543 | Manufacture of cocoa, chocolate and sugar confectionery | 1 |
| 311340 | Non-Chocolate Confectionery Manufacturing | D1543 | Manufacture of cocoa, chocolate and sugar confectionery | 1 |
| 212210 | Iron Ore Mining | C1310 | Mining of iron ores | 1 |
Now all that remains is to prepare the split links. Similar to the
disaggregation example above, we need to design weights to allocate the
“pool” of numeric value associated with the NAICS class
483213 into the corresponding ISIC classes
I6110 and I6120.
In this third example, one souce NAICS Canada class is equivalent to more than one target ISIC class, forming a one-to-many relation.
canada_split <- tibble::tribble(
~NAICS1997, ~NAICS1997_desc, ~ISIC3, ~ISIC3_desc, ~Link,
"483213", "Inland Water Transportation (except by Ferries)", "I6110 *", "Sea and coastal water transport", "Intracoastal water transportation",
"483213", "Inland Water Transportation (except by Ferries)", "I6120 *", "Inland water transport", "Inland water transportation (except ferries)"
)| NAICS1997 | NAICS1997_desc | ISIC3 | ISIC3_desc | Link |
|---|---|---|---|---|
| 483213 | Inland Water Transportation (except by Ferries) | I6110 * | Sea and coastal water transport | Intracoastal water transportation |
| 483213 | Inland Water Transportation (except by Ferries) | I6120 * | Inland water transport | Inland water transportation (except ferries) |
Assume for illustration purposes that we have reference data to
suggest the Canadian “Inland water transport” industry
(I6120) is twice as big as the “Sea and coastal water
transport” industry (I6110). This suggests that the weight
between 483213 and I6120 should be twice that
of I6110.
canada_frac <- canada_split |>
dplyr::mutate(ISIC3 = stringr::str_remove(ISIC3, " \\*")) |>
dplyr::select(-Link) |>
dplyr::mutate(share = dplyr::case_when(
ISIC3 == "I6110" ~ 0.33,
ISIC3 == "I6120" ~ 0.67,
T ~ NA_real_
))Now let’s combine the unit and fractional links into a crossmap:
canada_xmap <- dplyr::bind_rows(canada_unit, canada_frac) |>
as_xmap_df(from = NAICS1997, to = ISIC3, weights = share)## Dropping any additional columns in `dplyr::bind_rows(canada_unit, canada_frac)`
## ℹ To silence set `.drop_extra = TRUE`
print(canada_xmap)## xmap_df:
## recode, split, and collapse
## (NAICS1997 -> ISIC3) BY share
## NAICS1997 ISIC3 share
## 1 311320 D1543 1.00
## 2 311330 D1543 1.00
## 3 311340 D1543 1.00
## 4 212210 C1310 1.00
## 5 483213 I6110 0.33
## 6 483213 I6120 0.67Reversing Crossmaps
One-Way Maps
Except in the case of recoding, crossmaps are generally lateral
(one-way). Weights on collapse and split links are no longer valid if
you reverse the direct of the link. Notice that
as_xmap_df() throws an error if you try to naively swap the
from and to arguments:
dplyr::bind_rows(canada_unit, canada_frac) |>
as_xmap_df(from = ISIC3, to = NAICS1997, weights = share)## Dropping any additional columns in `dplyr::bind_rows(canada_unit, canada_frac)`
## ℹ To silence set `.drop_extra = TRUE`## Error in `as_xmap_df()`:
## ! Incomplete mapping weights found
## ✖ `share` does not sum to 1
## ℹ Modify weights or adjust `tol` and try again.Reversible Maps
However, we can swap the arguments on a recode map. Recall the ISO country code crossmap we created above:
print(iso_xmap)## xmap_df:
## recode
## (ISONumeric -> ISO2) BY weight
## ISONumeric ISO2 weight
## 1 004 AF 1
## 2 008 AL 1
## 3 012 DZ 1
## 4 016 AS 1
## 5 020 AD 1Imagine that instead of converting country codes from ISO Numeric to ISO-2 digit, we wanted to convert from ISO-2 digit to ISO Numeric. We can take the existing crossmap and invert it without editing any weights:
iso_xmap |>
xmap_reverse()## xmap_df:
## recode
## (ISO2 -> ISONumeric) BY r_weights
## ISO2 ISONumeric r_weights
## 1 AF 004 1
## 2 AL 008 1
## 3 DZ 012 1
## 4 AS 016 1
## 5 AD 020 1A less trivial reversal is creating an aggregation map from a disaggregation map. Recall the country to state level disaggregation map we created above:
state_xmap |>
xmap_drop_extra()## xmap_df:
## split
## (ctr -> adm1) BY share
## ctr adm1 share
## 1 AU AU-NSW 0.31391391
## 2 AU AU-VIC 0.25462770
## 3 AU AU-QLD 0.20490105
## 4 AU AU-SA 0.07008932
## 5 AU AU-WA 0.10723416
## 6 AU AU-TAS 0.02200277
## 7 AU AU-NT 0.00964811
## 8 AU AU-ACT 0.01758297Now imagine we wanted to re-aggregate the data, say after some state-level adjustments:
state_xmap |>
xmap_reverse(weights_into = "agg_w") |>
xmap_drop_extra()## xmap_df:
## recode and collapse
## (adm1 -> ctr) BY agg_w
## adm1 ctr agg_w
## 1 AU-NSW AU 1
## 2 AU-VIC AU 1
## 3 AU-QLD AU 1
## 4 AU-SA AU 1
## 5 AU-WA AU 1
## 6 AU-TAS AU 1
## 7 AU-NT AU 1
## 8 AU-ACT AU 1